Bootstrap AUC: Difference between revisions
Jump to navigation
Jump to search
| Line 24: | Line 24: | ||
[[File:fig_compare_methods.png|thumb|center|375px]] | [[File:fig_compare_methods.png|thumb|center|375px]] | ||
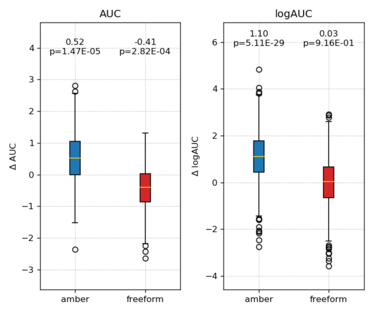
Delta AUC and delta logAUC will be computed and displayed. | |||
The p-value from paired t-test indicate if change is statistically significant or not. | |||
Revision as of 23:18, 25 February 2019
To test whether the difference in AUC/logAUC between two methods is statistically significant or not, AUC/logAUC of the new developed method(s) against the reference method can be compared with bootstrap.
Files needed:
- ligands.name --> file with ligand names to perform enrichment
- decoys.name --> file with decoy names to perform enrichment
- score file(s) --> extract_all.sort.uniq.txt
First, the anaconda python environment needs to be set:
source /nfs/home/yingyang/.cshrc_anaconda
Plot the variation of AUC/logAUC
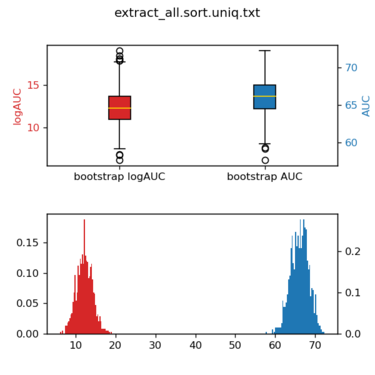
python /nfs/home/yingyang/work/scripts/bootstrap_AUC.py \ -l ./ligands.name -d ./decoys.name \ -s1 extract_all.sort.uniq.txt \ -p single
The figure will looks like this:
Plot the change(s) in AUC/logAUC against reference score
python /nfs/home/yingyang/work/scripts/bootstrap_AUC.py \ -l ../ligands.name -d ../decoys.name \ -s1 score.standard -s2 score.amber score.freeform \ -p compare
Delta AUC and delta logAUC will be computed and displayed. The p-value from paired t-test indicate if change is statistically significant or not.