Filtering ligands for novelty: Difference between revisions
Jump to navigation
Jump to search
Chasemwebb (talk | contribs) No edit summary |
Chasemwebb (talk | contribs) No edit summary |
||
| Line 1: | Line 1: | ||
Written by Chase Webb 09-01-2018 | Written by Chase Webb 09-01-2018 | ||
After a large scale docking campaign, it is important to remove prospective ligands that are too similar to compounds that are already known to modulate the receptor. In this way, we can focus on assessing new chemical interactions. | After a large scale docking campaign, it is important to remove prospective ligands that are too similar to compounds that are already known to modulate the receptor. In this way, we can focus on assessing new chemical interactions. This is best completed after clustering has been conducted as specified here [ | ||
=This process proceeds in the following steps:= | =This process proceeds in the following steps:= | ||
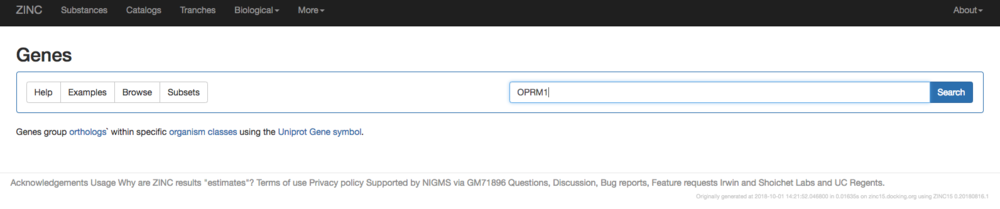
1. '''Generate a list of smiles for the known compounds.''' The most simple way to do this is to download them from ZINC. For the Mu opioid receptor (OPRM1) for instance, go here: [https://zinc15.docking.org/genes/home/ ZINC15 Genes] | 1. '''Generate a list of smiles for the known compounds.''' The most simple way to do this is to download them from ZINC. For the Mu opioid receptor (OPRM1) for instance, go here: [https://zinc15.docking.org/genes/home/ ZINC15 Genes] | ||
[[File:filter_ligands_image1.png|thumb|center| | [[File:filter_ligands_image1.png|thumb|center|1000px|Search for Your Molecules in ZINC Using the UNIPROT Ascension ID for Your Target, for example OPRM1 for the Mu Receptor]] | ||
2. '''Generate Fingerprints for the known compounds''' Run the following script written by TEB and JKL. The inputs are name of the knowns file and the name of the output fingerprint file. | |||
python ~jklyu/zzz.github/ChemInfTools/utils/teb_chemaxon_cheminf_tools/generate_chemaxon_fingerprints.py knowns_list.smi knowns | |||
3. | |||
Revision as of 21:56, 1 October 2018
Written by Chase Webb 09-01-2018
After a large scale docking campaign, it is important to remove prospective ligands that are too similar to compounds that are already known to modulate the receptor. In this way, we can focus on assessing new chemical interactions. This is best completed after clustering has been conducted as specified here [
This process proceeds in the following steps:
1. Generate a list of smiles for the known compounds. The most simple way to do this is to download them from ZINC. For the Mu opioid receptor (OPRM1) for instance, go here: ZINC15 Genes
2. Generate Fingerprints for the known compounds Run the following script written by TEB and JKL. The inputs are name of the knowns file and the name of the output fingerprint file.
python ~jklyu/zzz.github/ChemInfTools/utils/teb_chemaxon_cheminf_tools/generate_chemaxon_fingerprints.py knowns_list.smi knowns
3.