Interaction Filtering: Difference between revisions
No edit summary |
No edit summary |
||
| Line 33: | Line 33: | ||
'''Column4: #unpaired ligand donor + #unpaired ligand salt bridge''' | '''Column4: #unpaired ligand donor + #unpaired ligand salt bridge''' | ||
'''Column9: # | '''Column9: #hydrogen bond''' | ||
'''Column11: #salt bridge''' | '''Column11: #salt bridge''' | ||
Revision as of 08:11, 17 June 2020
This is the 1st version (20200210) of Interaction Filtering. Please copy the code to your current directory.
$ cp -r /mnt/nfs/home/sgu/code/interfilter .
FIRST AND FOREMOST, if you find any version conflicts between python3 and python2, you need to comment out inside ~/.cshrc: # source /nfs/soft/dock/versions/dock37/DOCK-3.7-trunk/env.csh
To run the code, you need to install OpenEye (version 2019.Oct.2) by following the instruction: https://docs.eyesopen.com/toolkits/python/quickstart-python/linuxosx.html
On our cluster, you may source my environment.
$ source /nfs/home/sgu/anaconda3/etc/profile.d/conda.csh $ conda activate oepython $ source /nfs/soft/openeye/license.csh
Running the code:
$ python interfilter.py -protein rec.crg.pdb -ligand poses.mol2
If you want to have/avoid the interaction (hydrogen bond or salt bridge) for a specific residue (e.g. ASP115A, A means chain A):
In rec.crg.pdb, some residue like HIS is converted to HID or HIE. Please use the converted name instead of HIS.
$ python interfilter.py -protein rec.crg.pdb -ligand poses.mol2 -residue ASP115A
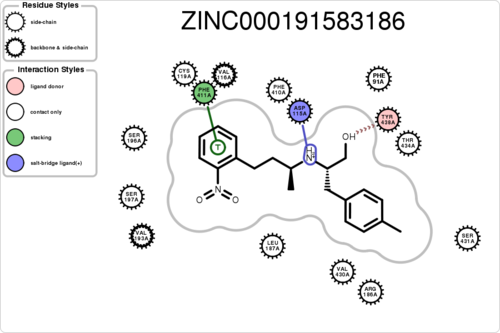
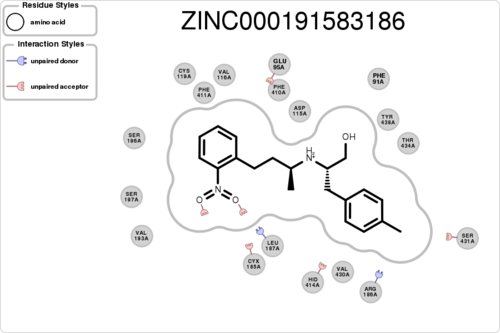
If you want to plot the paired/unpaired interaction (figure generation can be slow, but tens of thousands should be fine):
$ python interfilter.py -protein rec.crg.pdb -ligand poses.mol2 -residue ASP115A -plot
The output is a txt file, containing 15 columns: 1ligand 2clash 3hbond_clash+sbridge_clash 4unpairedl_donor+unpairedl_sbridge 5unpairedl_acceptor 6unpairedp_donor+unpairedp_acceptor 7unpairedp_sbridge 8contact 9hbond+hbond_nonideal 10hbond_ligand 11sbridge 12stacking 13cationpi 14halogen 15residue
To filter out compounds, you may be interested in:
Column3: #hydrogen bond clash + #salt bridge clash
Column4: #unpaired ligand donor + #unpaired ligand salt bridge
Column9: #hydrogen bond
Column11: #salt bridge
Column15: interaction (hydrogen bond or salt bridge) for you specified residue (0 means no, 1 means yes)
$ awk '$3==0 && $4==0 && $9+$11>0' poses_noResidue_interaction_analysis.txt > filtered.txt $ awk '$3==0 && $4==0 && $9+$11>0 && $15>0' poses_ASP115A_interaction_analysis.txt > filtered.txt