DOCK Blaster:Tutorial 2: Difference between revisions
No edit summary |
No edit summary |
||
| Line 71: | Line 71: | ||
== Possible problems and alternative approaches == | == Possible problems and alternative approaches == | ||
[[Category:DOCK Blaster]] | |||
[[Category:Tutorials]] | |||
Revision as of 18:20, 21 September 2007
DOCK Blaster Tutorial 1. Docking Methotrexate (MTX) to Dhihydrofolate Reductase (DHFR)
This is one of the DOCK Blaster:Tutorials.
Conceptual Overview
Methotrexate (MTX, depicted) binds to human dihydrofolate reductase (DHFR) with high affinity, and has been used in clinical oncology for over 20 years. A crystal structure is available (PDB code 3dfr, depicted) with MTX in the binding site.
- Question 1. Is DOCK Blaster able to suggest other compounds from a commercially available library that might be active against dihydrofolate reductase?
- Sub-question 1.1. Is DOCK Blaster able to re-dock MTX close to its crystallographically observed position, with a competitive score?
- Sub-question 1.2. Is DOCK Blaster able to enrich MTX compared with 50 property-matched decoys, compounds with similar properties to MTX but dissimilar topology?
- Sub-question 1.3. Is DOCK Blaster able to enrich known actives, and de-enrich known inactives, against DHFR?
picture of MTX
picture of 3dfr
picture of MTX bound to 3dfr
picture of other DHFR ligands
From Theory to Action
We will attempt to use DOCK Blaster to answer these questions, using data that has been prepared in advance. If you wish to use DOCK Blaster on your own project, you must prepare data to conform with DOCK Blaster's basic requirements. To do this tutorial, please carefully follow the steps below. This tutorial usually takes about two hours, but most of the time is simply waiting for the computers to do the work on our servers.
Literature References
McMaster study Blaney's Magnum opus.
Collecting and formatting relevant data
Launch Preliminary Calculation
- 1. Go to DOCK Blaster web page blaster.docking.org. Please note that during pre-alpha testing this page is different. Contact John Irwin for details.
- 2. Click on "Start docking!" to go to the input preparation page.
- 3. Click on "HELP" to open up the documentation for input preparation in a separate window.
- 3.1. In the documentation window, click on "Sample Data" at the end of the first paragraph.
- 3.2 In the sample data window, click on the link to data.docking.org
- 3.3. Click on DUD40.
- 3.4. Click on CDK2.
- 3.5. Right mouse to save all five files in this directory on your disk.
- 3.6. Close the documentation window, returning to the DOCK Blaster input form.
- 4. Click on the first "Browse" button (to the right of Target) and select the rec.pdb file you just downloaded.
- 5. In the next field "docked ligand" select the xtal-lig.mol2 file.
- 6. Optionally, you may select the actives.smi and inactives.smi from the next two lines.
- 7. Enter your email address if you wish to receive progress reports by email.
- 8. Enter a brief comment about the calculation.
- 9. (optional: enter cofactor.par below)
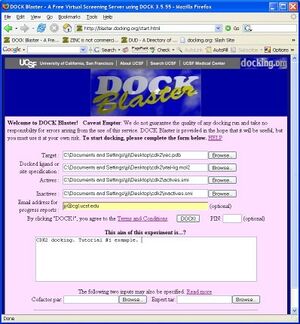
If you have followed the steps above, your screen should now look something like Figure Tut1-1 (right).
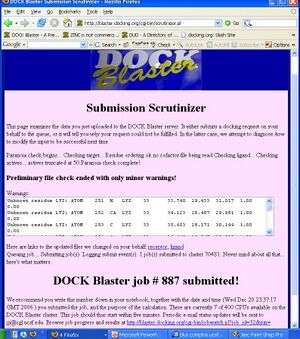
- 10. Click on "DOCK" to upload the files and begin docking. You will be taken to the "Submission Scrutinizer", as depicted in Figure Tut1-2.
- 11. Your job should now be running. If it is not, there is either some part of the above instructions you did not follow, or we are currently experiencing problems with our systems.
- 12. Read through the submission scrutinizer, and go to the Job Watcher by clicking on the link at the bottom of the page... http://blaster.docking.org/cgi-bin/jobmon.pl
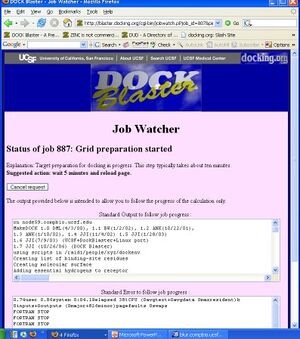
- 13. You should now be in the Job Watcher. Depending on how much time has elapsed, and how busy our cluster is, you will see a progress report of your job. It might look something like Figure Tut1-3.
- 14. If all goes well, the job will complete in about an hour. You may reload the page at anytime to get updated status of the job progress.
- 15. Preliminary docking normally begins after the site preparation is complete, often about 15 minutes after you first submitted the job. Once preliminary docking results are available, new links appear in the job watcher to allow you to start to see docked ligands. Caution: until the preliminary docking is complete, it can be misleading to rely upon incomplete preliminary docking results to reach any conclusions about the docking results.