DOCK Blaster:Tutorial 1: Difference between revisions
No edit summary |
No edit summary |
||
| Line 23: | Line 23: | ||
We will attempt to use DOCK Blaster to answer these questions, using data that has been prepared in advance. If you wish to use DOCK Blaster on your own project, you must prepare the data yourself. See [[DOCK Blaster:Input Preparation | Input Preparation]] for details. | We will attempt to use DOCK Blaster to answer these questions, using data that has been prepared in advance. If you wish to use DOCK Blaster on your own project, you must prepare the data yourself. See [[DOCK Blaster:Input Preparation | Input Preparation]] for details. | ||
== Protocol P1 - | == Protocol P1 - Prepare input files == | ||
* P1.1. Go to the benchmarking data page [http://data.docking.org/ data.docking.org] and browse the DUD40 set, selecting the [http://data.docking.org/dud40/mr/ MR set]. Download each of these five files. On windows, you do this by right mouse clicking and selecting "save link as". If you are using your own target, you must prepare the files yourself (see [[DOCK Blaster:Input Preparation | Input Preparation]]). | * P1.1. Go to the benchmarking data page [http://data.docking.org/ data.docking.org] and browse the DUD40 set, selecting the [http://data.docking.org/dud40/mr/ MR set]. Download each of these five files. On windows, you do this by right mouse clicking and selecting "save link as". If you are using your own target, you must prepare the files yourself (see [[DOCK Blaster:Input Preparation | Input Preparation]]). | ||
[[Image:DBtut1_1f.jpg|thumb|right|Figure T1.1 - Prepare input and launch job]] | [[Image:DBtut1_1f.jpg|thumb|right|Figure T1.1 - Prepare input and launch job]] | ||
== Protocol P2 - Submit job and review | == Protocol P2 - Submit job and review results == | ||
* P2.1. Go to the [http://blaster.docking.org | DOCK Blaster main page] and select [http://blaster.docking.org/start.shtml | Queue A Job] from the DOCK! pull down menu (Figure T1.1). Fill in the form as in the figure, as follows. | * P2.1. Go to the [http://blaster.docking.org | DOCK Blaster main page] and select [http://blaster.docking.org/start.shtml | Queue A Job] from the DOCK! pull down menu (Figure T1.1). Fill in the form as in the figure, as follows. | ||
** a. In the "Target" field, select the receptor, rec.pdb. | ** a. In the "Target" field, select the receptor, rec.pdb. | ||
| Line 57: | Line 57: | ||
Preliminary results above show that docking is able to enrich known actives from among a database of decoys. This suggests that prospective docking for discovery of novel ligands is reasonable and can be expected to suggest interesting compounds to test. | Preliminary results above show that docking is able to enrich known actives from among a database of decoys. This suggests that prospective docking for discovery of novel ligands is reasonable and can be expected to suggest interesting compounds to test. | ||
== Protocol P3. Dock | == Protocol P3. Dock purchasable fragments == | ||
== Protocol P4. Review and interpret | == Protocol P4. Review and interpret docking == | ||
== Protocol P5. | == Protocol P5. Prioritize for testing == | ||
== Possible problems | == Protocol P6. Suggested variations == | ||
== Possible problems & other approaches == | |||
= Literature Cited = | = Literature Cited = | ||
Revision as of 18:59, 27 November 2007
DOCK Blaster Tutorial 1. Dock to Mineralocorticoid receptor A DOCK Blaster Tutorial.
Specific Aims
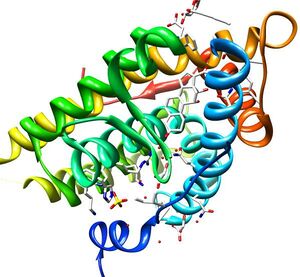
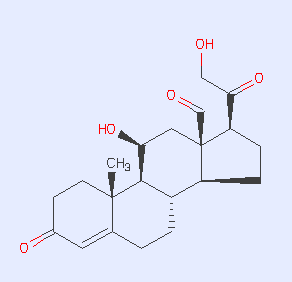
This tutorial will show you how to retrospectively dock a crystallographically observed ligand, aldosterone, into the human mineralocorticoid receptor, PDB code 2aa2. You will also dock a list of annotated actives from the literature, as well as property-matched decoys. Finally, you will prospectively dock the ZINC "fragment-like" library into MR, and pick compounds to test as possible binders. The questions are:
- 1. Can DOCK Blaster re-dock the native ligand close to its crystallographically observed position, with a competitive score?
- 2. Can DOCK Blaster enrich known actives from a database of property-matched decoys?
- 3. Can DOCK Blaster suggest novel, commercially available ligands for MR?
Background and Significance
Mineralocorticoid receptor (MR) is a receptor with high affinity for mineralocorticoids. It belongs to the steroid hormone receptor family where the ligand diffuses into cells, interacts with the receptor and results in a single transduction affecting specific gene expression in the nucleus. see Wikipedia.
Hippocampal mineralocorticoid receptors play a major role in the control of the hypothalamus-pituitary-adrena (HPA) axis, and thus MR is a target for drug discovery.
Preliminary Results
We will attempt to use DOCK Blaster to answer these questions, using data that has been prepared in advance. If you wish to use DOCK Blaster on your own project, you must prepare the data yourself. See Input Preparation for details.
Protocol P1 - Prepare input files
- P1.1. Go to the benchmarking data page data.docking.org and browse the DUD40 set, selecting the MR set. Download each of these five files. On windows, you do this by right mouse clicking and selecting "save link as". If you are using your own target, you must prepare the files yourself (see Input Preparation).
Protocol P2 - Submit job and review results
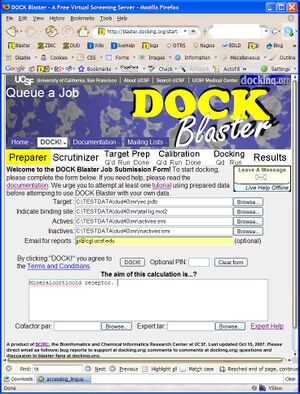
- P2.1. Go to the | DOCK Blaster main page and select | Queue A Job from the DOCK! pull down menu (Figure T1.1). Fill in the form as in the figure, as follows.
- a. In the "Target" field, select the receptor, rec.pdb.
- b. In the next field, select "Docked ligand" and select the ligand, xtal-lig.mol2.
- c. In the "Actives" field, select the actives.smi file.
- d. In the "Inactives" field, select the inactives.smi file.
- e. In "Email for reports", enter your email address (optional).
- f. In the "Aim of this experiment" field, write a short memo about what you are doing.
- g. Check your input, and click "DOCK!" when you are ready.
If you have followed the steps above, your screen should now look something like Figure Tut1-1 (right). [[Image:DBtut1_2f.jpg|thumb|right|Figure T1.2 - Scrutinizer - data are uploaded, checked, and the job is started].
- P2.2. Depending on available system resources, your job is now either running, or queued to run when a slot becomes available. If your screen does not look like Figure T1.2, check the input and try again. If the job is still failing after the third attempt, please write to support at docking.org for assistance.
- P2.3. Read what is reported by the Scrutinizer (Figure T1.2), and proceed to the Job Watcher by clicking on the link at the bottom of the page when you are ready.
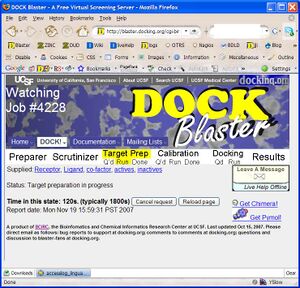
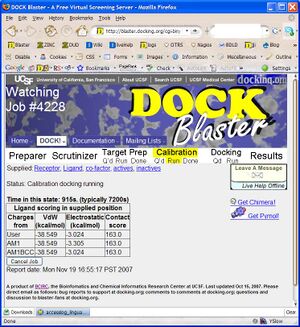
- P2.4. You should now be in the Job Watcher (Figure T1.3). Depending on how much time has elapsed, and how busy our cluster is, you will see a progress report of the current status of your job. As time passes, the yellow status indicator should progress from left to right indicating the progress of your job through Target Preparation and Calibration.
- P2.5. If all goes as usual, your job will have completed within an hour. You may reload the Job Watcher at any time to get an updated status report of your job. When calibration is complete, you may review the preliminary results and decide on the next course of action.
- P2.6 Review Calibration results. blah blah blah.
Proposed Research
Preliminary results above show that docking is able to enrich known actives from among a database of decoys. This suggests that prospective docking for discovery of novel ligands is reasonable and can be expected to suggest interesting compounds to test.
Protocol P3. Dock purchasable fragments
Protocol P4. Review and interpret docking
Protocol P5. Prioritize for testing
Protocol P6. Suggested variations
Possible problems & other approaches
Literature Cited
We refer you to our publications pages for theory and practice of docking and virtual screening.
- Publications:DOCK
- Publications:ZINC
- Publications:DUD
- Publications:Reviews_and_opinion
- Publications:Targets
Document status: working tutorial. Please correct any errors you may find.