AutoQSAR/DeepChem for billions of molecules: Difference between revisions
No edit summary |
No edit summary |
||
| (One intermediate revision by the same user not shown) | |||
| Line 56: | Line 56: | ||
scp -rp 1/ dt2.wynton.ucsf.edu:<path_to_where_to_run_prediction> | scp -rp 1/ dt2.wynton.ucsf.edu:<path_to_where_to_run_prediction> | ||
== Apply the ML model to predict all molecules (smiles) of interest | == Apply the ML model to predict all molecules (smiles) of interest == | ||
Model prediction can leverage large number of CPUs thus will be on wynton | |||
=== Prepare molecules (smiles) for prediction === | === Prepare molecules (smiles) for prediction === | ||
| Line 111: | Line 112: | ||
Once complete, check if the number of files in the standardize folder is the same as in the raw folder | Once complete, check if the number of files in the standardize folder is the same as in the raw folder | ||
=== Run prediction === | === Run ML prediction === | ||
Go to the folder where to run prediction (where ML model (1/) is located). | Go to the folder where to run prediction (where ML model (1/) is located). | ||
mkdir prediction | mkdir prediction | ||
Latest revision as of 19:12, 12 March 2021
2/25/2021 Ying Yang
Train a ligand-based ML model
Model training requires GPU thus will be on gimel5
- Prepare the input file for training in the format of <ligand smiles>,<dock score>
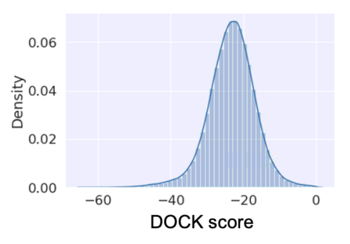
SMILES,DOCK score C[C@H]1COCCCN1C(=O)[C@@H]1CN2CCN1CCC2,-15.98 Cc1ccccc1-c1cc(C(=O)N2CC3(CN(C)C3)C2)n[nH]1,-17.43 CNC(=O)c1ccccc1NC(=O)[C@H]1C[C@H]2CCCCN2C1,-21.03 CC[C@H](F)CN[C@@H](CNC(=O)c1ccc(F)cc1F)C(C)C,4.73 C[C@@H]1CN(C(=O)C(=O)N[C@@H](c2cccc(F)c2)c2ccccn2)C[C@H]1N,13.34 CC(C)[C@H](NC(=O)NC[C@@H]1CCN(CCc2ccccc2)C1)C1CC1,5.9 CC[C@@H](F)CNCC1CCN(C(=O)c2ccc(F)s2)CC1,-14.68 Cc1cccc(C[C@@H](NCc2cccn2C)C2CC2)c1,-40.38 CC(C)NC(=O)c1ccc2nnc([C@H]3CN(Cc4ccccc4)CCN3C)n2c1,-23.15
IMHO, ML algorithm performs well with a normal distribution.
If < 30% molecules cannot be docked, it's safe to ignore the non-dockable (those without a dock score).
- Prepare the submission file, and submit to
cat <<EOF > sbatch_ml_train.sh
#!/bin/bash
#SBATCH --job-name=ml_deepchem
#SBATCH --partition=gimel5.gpu
#SBATCH --gres=gpu:1
#SBATCH --ntasks=1
hostname
nvidia-smi
source /nfs/home/yingyang/programs/ligand_ml/anaconda/etc/profile.d/conda.sh
conda activate ligand_ml
# CHANGE here: to your prepared input file
infile=AL-dock_5HT5a_train.csv
i=1
ligand_ml train model_\${i} \${infile}
ligand_ml package model_\${i} \${i}
EOF
sbatch sbatch_ml_train.sh
Once the model training complete, you will see a folder 1/ and a file 1.tar.gz.
Transfer the ML model in folder 1/ to Wynton
scp -rp 1/ dt2.wynton.ucsf.edu:<path_to_where_to_run_prediction>
Apply the ML model to predict all molecules (smiles) of interest
Model prediction can leverage large number of CPUs thus will be on wynton
Prepare molecules (smiles) for prediction
For example, H26.smi is the file including ZINC ids and smiles of molecules for prediction
- set up the folder and break down the input smiles
mkdir raw cd raw split -l 50000 ../H26.smi - cd ../
- run molecules standardization
mkdir -p standardized
set num=` ls raw/* | wc -l `
echo "Number of file to process:" $num
cat << EOF > qsub_standardize.csh
#\$ -S /bin/csh
#\$ -cwd
#\$ -pe smp 1
#\$ -l mem_free=2G
#\$ -l scratch=5G
#\$ -l h_rt=02:00:00
#\$ -j yes
#\$ -o std.out
#\$ -t 1-$num
hostname
date
echo ""
source /wynton/home/shoichetlab/yingyang/programs/ligand_ml/anaconda/etc/profile.d/conda.csh
conda activate ligand_ml
which python
which ligand_ml
echo ""
set BASE_DIR=\`pwd\`
# Do Not Edit Below This Point
set RAW_DIR=\${BASE_DIR}/raw/
set DEST_DIR=\${BASE_DIR}/standardized/
set CUDA_VISIBLE_DEVICES=-1
python /wynton/home/shoichetlab/yingyang/scripts_ML/code/standardize.py \$SGE_TASK_ID \${RAW_DIR} \${DEST_DIR}
EOF
qsub qsub_standardize.csh
Once complete, check if the number of files in the standardize folder is the same as in the raw folder
Run ML prediction
Go to the folder where to run prediction (where ML model (1/) is located).
mkdir prediction
set in=./
set model=1
set out=prediction_1
mkdir -p ${out}
set num=` ls standardized/*.csv | wc -l `
echo "Number of file to process:" $num
cat << EOF >! qsub_ml_${hac}.sh
#\$ -S /bin/sh
#\$ -cwd
#\$ -pe smp 1
#\$ -l mem_free=50G
#\$ -l scratch=50G
#\$ -l h_rt=01:00:00
#\$ -o qsub_ml_${hac}.out
#\$ -j yes
#\$ -t 1-$num
hostname
date
echo ""
source /wynton/home/shoichetlab/yingyang/programs/ligand_ml/anaconda/etc/profile.d/conda.sh
conda activate ligand_ml
which python
which ligand_ml
echo ""
# Set your variables
export TRAIN_DIR=\$(pwd)/${in}
export MODEL_NAME=$model
export MODEL=\${TRAIN_DIR}/\${MODEL_NAME}
export BASE_DIR=\$(pwd)/
export DEST_DIR=\${BASE_DIR}/${out}/
# Do Not Edit Below This Point
export CODE_DIR=/wynton/home/shoichetlab/yingyang/scripts_ML/code
export STANDARDIZED_DIR=\${BASE_DIR}/standardized/
export INFILE=\${STANDARDIZED_DIR}/\${SGE_TASK_ID}.csv
export OUTFILE=\${DEST_DIR}/\${SGE_TASK_ID}.csv
# Do path magic to set things up and use 1 cpu
#export LD_LIBRARY_PATH=/nfs/soft/schrodinger/2019-4/internal/lib/cuda-stubs/:\$LD_LIBRARY_PATH
export LD_LIBRARY_PATH=/wynton/home/shoichetlab/yingyang/cuda-stubs/:\$LD_LIBRARY_PATH
export CUDA_VISIBLE_DEVICES=-1
export CPU_STATS=\$(cat /dev/urandom | tr -cd 'a-f0-9' | head -c 32)
export MY_CPU_FILE=\$(cat /dev/urandom | tr -cd 'a-f0-9' | head -c 32)
sleep \$[ ( \$RANDOM % 10 ) ]s
mpstat -P ALL 5 1 > /tmp/\$CPU_STATS
python \${CODE_DIR}/get_idle_cpu.py /tmp/\$CPU_STATS /tmp/\$MY_CPU_FILE
export MY_CPU=\$(cat /tmp/\$MY_CPU_FILE)
echo "Using cpu \$MY_CPU"
n=0
until [ \$n -ge 5 ]
do
echo "taskset -c \$MY_CPU ligand_ml evaluate \$MODEL \$INFILE \$OUTFILE --skip_standardization=True --skip_version_check=True"
taskset -c \$MY_CPU ligand_ml evaluate \$MODEL \$INFILE \$OUTFILE --skip_standardization=True --skip_version_check=True && break
n=\$[\$n+1]
sleep 5
done
EOF
qsub qsub_ml.sh
Analyze prediction
Get the top 5% of ML prediction. A larger memory node is recommended for sorting...
source /wynton/home/shoichetlab/yingyang/programs/miniconda3/etc/profile.d/conda.sh
conda activate opencadd
# 50,000 mols per file
# 12,500 --> 25%
# 25,000 --> 5%
# 5,000 --> 1%
# CHANGE here: to the prediction folder
dir_ml=prediction_1
echo "Process ${dir_ml} ... "
rm ml_5percent.csv
touch ml_5percent.csv
for f in $(ls ${dir_ml}/* ); do
echo $f
head -n 25000 ${f} | egrep -hiv 'score|model' >> ml_5percent.csv
done
python /wynton/home/shoichetlab/yingyang/scripts_ML/out_analysis/sort_qsar_prediction.py ml_5percent.csv
rm ml_5percent.csv
Once complete, the top 5% from ML prediction will be in ml_5percent_sort.csv
The same procedure can be applied to other scores: dock scores, FEP predicted values...