Generating extrema set: Difference between revisions
No edit summary |
No edit summary |
||
| Line 42: | Line 42: | ||
4) Output files | 4) Output files | ||
4.1) (charge-type)_tranche_summary.txt | |||
4.1) (charge-type)_tranche_summary.txt. The file contains how many molecules has been selected from each tranche. The section below is an example: | |||
EF 430 | EF 430 | ||
DF 337 | DF 337 | ||
| Line 54: | Line 55: | ||
1994 | 1994 | ||
4.2) (charge-type)_charge_tranches.list | 4.2) (charge-type)_charge_tranches.list. The file contains the db2 index that has been selected from the extrema generation. | ||
5) Combine all the db2 indexes generated by the script. | |||
cat *_charge_tranches.list > extrema_set.list | |||
Then use the extrema_set.list as .sdi to set up your docking screen and run it. | |||
Revision as of 21:30, 12 October 2019
Written by Jiankun Lyu, 2019/10/12
extrema_set_gen------- working
|
|------ ZINC-downloader-3D-minu2.database_index
|
|------ ZINC-downloader-3D-minu1.database_index
|
|------ ZINC-downloader-3D-neutral.database_index
|
|------ ZINC-downloader-3D-plus1.database_index
|
|------ ZINC-downloader-3D-plus2.database_index
1) Make those directories above.
mkdir extrema_set_gen cd extrema_set_gen mkdir working
2) Download databases index from ZINC with different charge types
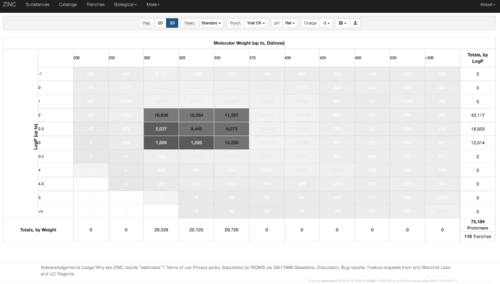
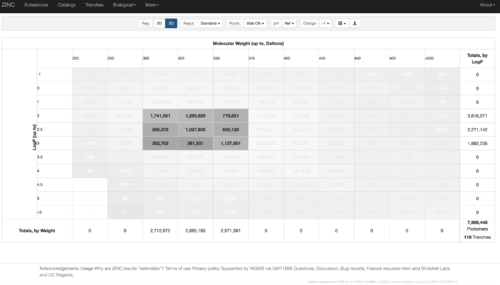
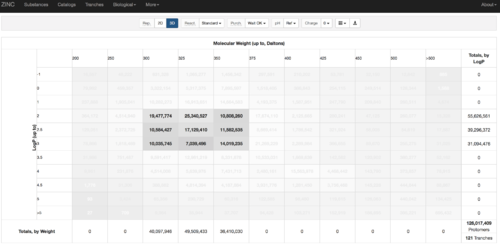
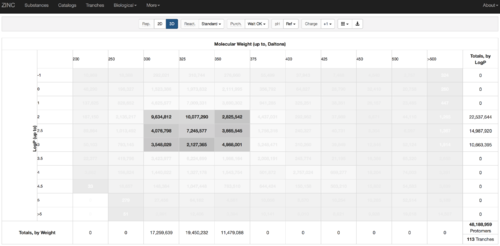
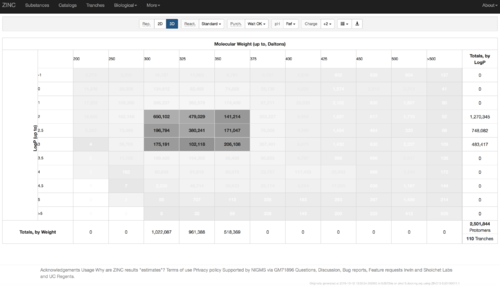
2.1) Go to ZINC http://zinc15.docking.org/tranches/home/#
2.2) Choose the tranches you want to generate extrema set for testing the charge preference. The goldilocks set has been chosen here as an example.
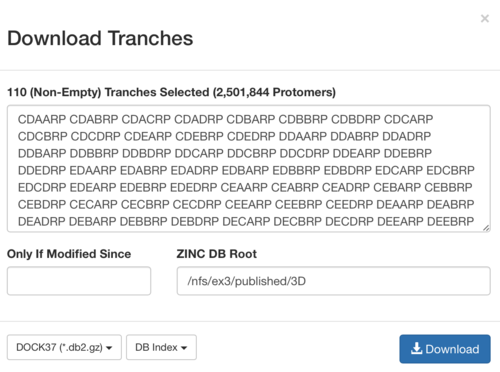
2.3) download the databases index file for each charge type
2.4) download the files above and save it as ZINC-downloader-3D-(charge-type).database_index, then upload the file to the working directory. In the working directory, you are supposed to have 5 files with names: ZINC-downloader-3D-minu2.database_index, ZINC-downloader-3D-minu1.database_index, ZINC-downloader-3D-neutral.database_index, ZINC-downloader-3D-plus1.database_index and ZINC-downloader-3D-plus2.database_index.
3) Run extrema set generation on 5 different charge types
python /mnt/nfs/ex5/work/jklyu/sigma2/gen_extrema/script/gen_extrema.py ZINC-downloader-3D-plus2.database_index 'plus2' 100 > log_plus2 & python /mnt/nfs/ex5/work/jklyu/sigma2/gen_extrema/script/gen_extrema.py ZINC-downloader-3D-plus1.database_index 'plus1' 100 > log_plus1 & python /mnt/nfs/ex5/work/jklyu/sigma2/gen_extrema/script/gen_extrema.py ZINC-downloader-3D-neutral.database_index '0' 100 > log_0 & python /mnt/nfs/ex5/work/jklyu/sigma2/gen_extrema/script/gen_extrema.py ZINC-downloader-3D-minus1.database_index 'minus1' 100 > log_minus1 & python /mnt/nfs/ex5/work/jklyu/sigma2/gen_extrema/script/gen_extrema.py ZINC-downloader-3D-minus2.database_index 'minus2' 100 > log_minus2 &
4) Output files
4.1) (charge-type)_tranche_summary.txt. The file contains how many molecules has been selected from each tranche. The section below is an example:
EF 430 DF 337 ED 293 DD 181 DE 120 CF 112 CE 131 CD 272 EE 118 1994
4.2) (charge-type)_charge_tranches.list. The file contains the db2 index that has been selected from the extrema generation.
5) Combine all the db2 indexes generated by the script.
cat *_charge_tranches.list > extrema_set.list
Then use the extrema_set.list as .sdi to set up your docking screen and run it.