FEP+ for GPCR: Difference between revisions
Chasemwebb (talk | contribs) No edit summary |
Chasemwebb (talk | contribs) No edit summary |
||
| Line 37: | Line 37: | ||
$SCHRODINGER/run membrane_cms2fep.py -ligand 'ligand' 2A_NBOH_MD-out.cms -o relax_2A_NBOH_pv.mae | $SCHRODINGER/run membrane_cms2fep.py -ligand 'ligand' 2A_NBOH_MD-out.cms -o relax_2A_NBOH_pv.mae | ||
You have now prepared the receptor and the initial state for an FEP+ map. | You have now prepared the receptor and the initial state for an FEP+ map. Marvel at your accomplishments. | ||
== Useful Schrodinger Commands == | == Useful Schrodinger Commands == | ||
| Line 47: | Line 47: | ||
== Ligand side == | == Ligand side == | ||
Careful preparation of the ligands is critical to a successful FEP+ prediction. Best practices include running LigPrep on all the compounds to exhaustively enumerate all the stereoisomers and likely protonation states of the ligands. Note that | Careful preparation of the ligands is critical to a successful FEP+ prediction. Best practices include running LigPrep on all the compounds to exhaustively enumerate all the stereoisomers and likely protonation states of the ligands. Note that tertiary amines cannot invert stereochemistry during the simulation, making it important to model both pseudo-stereoisomers. Note: This is especially important for basic amines that are protonated. Make sure to model the proton on both sides! | ||
* Import the ligands into the workspace, recommend that you import smiles or a mol2 file. Use the '''LigPrep''' tool to build the ligands into a 3D dockable format. Consider generating a good amount of stereoisomers if you are unsure about the sterochemistry or you only have relative stereochemical information. Generate the relevant protomers at the relevant pH. | |||
* Force field builder | * Force field builder | ||
Revision as of 03:09, 9 March 2021
2/25/2021 Ying Yang
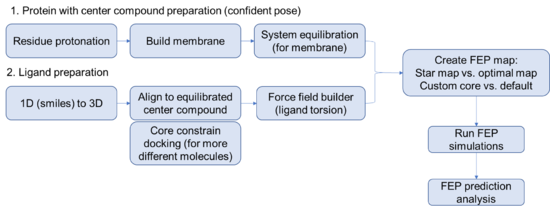
Steps for setting up a FEP prediction for membrane protein
Protein side
- Its always good practice to specify a relevant working directory and save the project. Maestro is prone to crashing randomly, especially when remotely accessing the license. Wouldn't want to lose all of your hard work.
- Begin by importing the structure of interest with a ligand that you are confident about. (FEP works best with an experimentally determined structure but can work with a docked pose if you are careful).
- Run the protein preparation wizard on your protein. Think carefully about protonation, you might even consider using the interactive protonation function to optimize the protonation state of the binding site residues or allosteric control points manually.
Protein model completeness Protein preparation should include fixing any chain breaks, modeling in any loop conformations and adding any missing side chains. Chain breaks near the active site will likely lead to poor results. Disulfide bridges should be created and termini residues capped where applicable.
Equilibration of complex structure (with confident binding pose)
- Build membrane using the System Builder tool. Use residue information obtained from OPM database (https://opm.phar.umich.edu/), add salts, add solvent. Default settings work well for most situations.
res.num 76-97,112-136,141,143,146-171,194-215,227-229,231-256,323-345,347,360-380,382,398
- Use the Molecular Dynamics tool to write out a simulation file to run on the cluster. Load the system you built in the previous step from the workspace. Specify a simulation time, for high confidence complexes, 20 ns should suffice. For more exploratory complexes and to assess stability of the ligand extend to 50-100 ns. Use the default Schrodinger protocol to relax the model system before starting the production run of the simulation. For ease of evaluation, you can check the box to output interaction analysis at the completion of the job. Instead of clicking run, click the gear to the left of the run button and write the file out to the disk.
- Replace the gimel-biggpu in the written out simulation job with the correct gpu location, gimel5.heavygpu then transfer (scp) the simulation job to gimel5, and submit
sed -i 's/gimel-biggpu/gimel5.heavygpu/g' desmond_md_job_1.sh bash desmond_md_job_1.sh
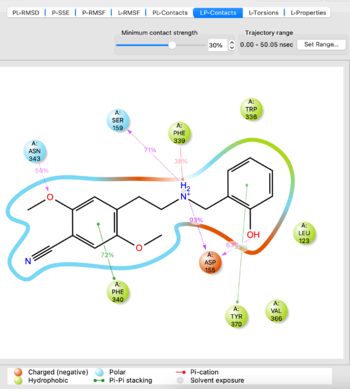
- Analyze the MD simulation using the Simulation Interactions Diagram tool from the TASKS menu. Import the -out.cms file from the simulation to analyze, it takes 5-10 minutes to load. If you are satisfied with the simulation as evidenced by things such as stability of protein/ligand, proceed. If not, potentially try another docked pose or run the simulation for a longer time.
Visualize the trajectory and analyze the simulation with SID tool
- Use the schrodinger command line tools to convert the last frame of the simulation -out.cms into .mae file.
$SCHRODINGER/run membrane_cms2fep.py -ligand 'ligand' 2A_NBOH_MD-out.cms -o relax_2A_NBOH_pv.mae
You have now prepared the receptor and the initial state for an FEP+ map. Marvel at your accomplishments.
Useful Schrodinger Commands
- Kill a submitted or running job:
$SCHRODINGER/jobcontrol -kill <jobID>
Ligand side
Careful preparation of the ligands is critical to a successful FEP+ prediction. Best practices include running LigPrep on all the compounds to exhaustively enumerate all the stereoisomers and likely protonation states of the ligands. Note that tertiary amines cannot invert stereochemistry during the simulation, making it important to model both pseudo-stereoisomers. Note: This is especially important for basic amines that are protonated. Make sure to model the proton on both sides!
- Import the ligands into the workspace, recommend that you import smiles or a mol2 file. Use the LigPrep tool to build the ligands into a 3D dockable format. Consider generating a good amount of stereoisomers if you are unsure about the sterochemistry or you only have relative stereochemical information. Generate the relevant protomers at the relevant pH.
- Force field builder
Run force field builder for all ligands
- Flexible ligand alignment OR core constrain docking
Depends on how similar/different are the ligands to the reference/center ligand
- Create FEP maps
- Write out the submission file; change host; submit on gimel5 via slurm