DOCK 3.7 tutorial based on Webinar 2017/06/28
Scenario 1: Use docking to predicted how Erlotinib (an approved drug) binds to the Epidermal Growth Factor Receptor
Get the link from the zinc webpage and us wget to download:
wget http://files.docking.org/protomers/16/89/55/209168955.db2.gz
Put the path of the downloaded database into the split database index file (this file usually contain many db2 file):
ls /path/tutorial_for_webinar/dock3.7/209168955.db2.gz > ligands.sdi
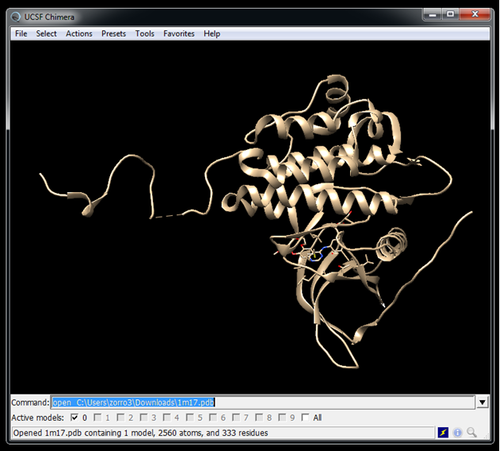
Get the receptor structure from the PDB website
wget https://files.rcsb.org/download/1M17.pdb --no-check-certificate
You may use a program like Chimera for this
Receptor file must be called: rec.pdb
Ligand file: xtal-lig.pdb
What if the crystal does not have a ligand:
Place atoms in the site were you want to dock. One way is to run sphgen and selecting spheres near residues in the site convert to pdb
Make the recptor file (remove alternative side chains):
grep "^ATOM" 1M17.pdb | grep -v ^................B > rec.pdb
Make ligand file:
grep AQ4 1M17.pdb | sed -e 's/HETATM/ATOM /g' > xtal-lig.pdb
Run blastermaster: input rec.pdb, xtal-lig.pdb and makes all receptor file need for docking.
python $DOCKBASE/proteins/blastermaster/blastermaster.py --addhOptions=" -HIS -FLIPs " -v
This command may take several minutes to run.